 
To do so, we created an initial word2vec model for immune system sequence embedding. Here, the described line of work is continued, with a focus on developing an embedding technique for B/T cell receptor HTS data, and using this embedding, along with machine learning abilities, to answer real-life questions in computational immunology. This approach opens the door for countless exciting opportunities for future developments. dna2vec provides experimental evidence that the arithmetic of the embedded vectors is akin to nucleotides’ concatenation, while ProtVec shows that tasks like protein family classification and disordered protein detection are not only feasible using the proposed representation and feature extraction method, but also outperform existing classification methods. These studies demonstrate the feasibility of sequence embedding. The main ones are ProtVec ( 8), seq2vec ( 9) and dna2vec ( 10), all use the word2vec concept introduced by ( 7). While there are countless different applications using the above methods, only few published works have implemented them for biological data analysis. In 2013, ( 7) brought word embedding to the fore by presenting the “word2vec” method, which is an NLP-embedding method based on an artificial neural networks, and became the basis for many of today’s NLP applications. Since then, vector space models for semantics are gradually developing and gaining popularity compared with traditional distributed representations. “Word embedding” was first introduced by ( 6). In NLP, the term “embedding” refers to the representation of symbolic information in text at the word-level, phrase-level, and even sentence-level, in terms of real number vectors. One possible approach to do so is to use existing embedding methods from the natural language processing (NLP) world, and adapt them to immunological sequences. The ability to embed these textual sequences in a vector-space is an important step towards developing effective analysis methods. The mathematical and statistical properties of high-dimensionality are often poorly understood or overlooked in data modeling and analysis ( 5). These technologies present investigators with the challenge of extracting meaningful statistical and biological information from high-dimensional data. With the advancements of HTS technologies, the amount of sequencing data is continuously growing ( 4). 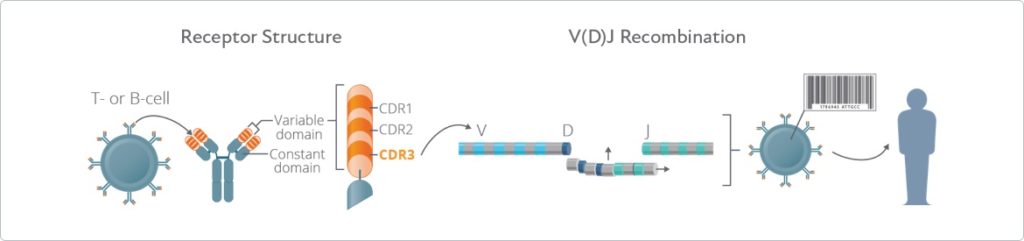
High Throughput Sequencing (HTS) is a powerful platform that enables large-scale characterization of BCR repertoires ( 3). The BCR repertoire in humans is estimated to include at least 10 11 different BCRs, and potentially several orders of magnitude greater ( 2). Applying Immune2vec along with machine learning approaches to patient data exemplifies how distinct clinical conditions can be effectively stratified, indicating that the embedding space can be used for feature extraction and exploratory data analysis.Īntibodies, the secreted form of BCRs, play a crucial role in the adaptive immune system, by binding specifically to pathogens and neutralizing their activity ( 1). Our work demonstrates Immune2vec to be a reliable low-dimensional representation that preserves relevant information of immune sequencing data, such as n-gram properties and IGHV gene family classification. We validate Immune2vec on amino acid 3-gram sequences, continuing to longer BCR sequences, and finally to entire repertoires. Here we present Immune2vec, an adaptation of a natural language processing (NLP)-based embedding technique for BCR repertoire sequencing data. The ability to embed these DNA and amino acid textual sequences in a vector-space is an important step towards developing effective analysis methods. These huge immune repertoires in each individual present investigators with the challenge of extracting meaningful biological information from multi-dimensional data. It does so through dynamic and diverse repertoires of T- and B- cell receptors (TCR and BCRs, respectively). The adaptive branch of the immune system learns pathogenic patterns and remembers them for future encounters. 2Bar Ilan Institute of Nanotechnologies and Advanced Materials, Bar Ilan University, Ramat Gan, Israel.1Bioengineering, Faculty of Engineering, Bar Ilan University, Ramat Gan, Israel.Miri Ostrovsky-Berman 1,2 Boaz Frankel 1,2 Pazit Polak 1,2 Gur Yaari 1,2* 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
